Today I’m going to walk through working with tags in Postgres with a sample database of 🐈 cats and their attributes
- First, I’ll look at a traditional relational model
- Second, I’ll look at using an integer array to store tags
- Lastly, I’ll test text arrays directly embedding the tags alongside the feline information
Tags in a relational model
For these tests, we will use a very simple table of 🐈 cats, our entity of
interest, and tags a short table of ten tags for the cats. In between the two
tables, the relationship between tags and cats is stored in
the cat_tags table.
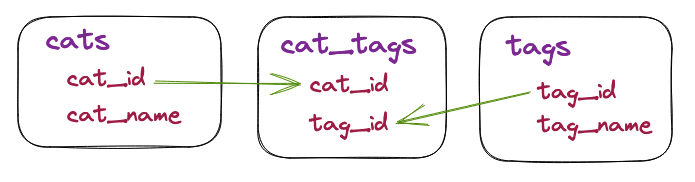
Table Creation SQL
CREATE TABLE cats (
cat_id serial primary key,
cat_name text not null
);
CREATE TABLE cat_tags (
cat_id integer not null,
tag_id integer not null,
unique(cat_id, tag_id)
);
CREATE TABLE tags (
tag_id serial primary key,
tag_name text not null,
unique(tag_name)
);
I filled the tables with over 1.7M entries for the cats, ten entries for
the tags, and 4.7M entries for the cat/tag relationship.
Data Generation SQL
-- Generate cats random names
INSERT INTO cats (cat_name)
WITH
hon AS (
SELECT *
FROM unnest(ARRAY['mr', 'ms', 'miss', 'doctor', 'frau', 'fraulein', 'missus', 'governer']) WITH ORDINALITY AS hon(n, i)
),
fn AS (
SELECT *
FROM unnest(ARRAY['flopsy', 'mopsey', 'whisper', 'fluffer', 'tigger', 'softly']) WITH ORDINALITY AS fn(n, i)
),
mn AS (
SELECT *
FROM unnest(ARRAY['biggles', 'wiggly', 'mossturn', 'leaflittle', 'flower', 'nonsuch']) WITH ORDINALITY AS mn(n, i)
),
ln AS (
SELECT *
FROM unnest(ARRAY['smithe-higgens', 'maclarter', 'ipswich', 'essex-howe', 'glumfort', 'pigeod']) WITH ORDINALITY AS ln(n, i)
)
SELECT initcap(concat_ws(' ', hon.n, fn.n, mn.n, ln.n)) AS name
FROM hon, fn, mn, ln, generate_series(1,5)
ORDER BY random();
-- Fill in the tag names
INSERT INTO tags (tag_name) VALUES
('soft'), ('cuddly'), ('brown'), ('red'), ('scratches'), ('hisses'), ('friendly'), ('aloof'), ('hungry'), ('birder'), ('mouser');
-- Generate random tagging. Every cat has 25% chance of getting each tag.
INSERT INTO cat_tags
WITH tag_ids AS (
SELECT DISTINCT tag_id FROM tags
),
tag_count AS (
SELECT Count(*) AS c FROM tags
)
SELECT cat_id, tag_id
FROM cats, tag_ids, tag_count
WHERE random() < 0.25;
— Create an index
CREATE INDEX cat_tags_x ON cat_tags (tag_id);
In total, the relational model needs 2424KB for the cats, tags and the tag
relationships.
SELECT pg_size_pretty(
pg_total_relation_size('cats') +
pg_total_relation_size('cat_tags') +
pg_total_relation_size('tags'));
Performance of relational queries
We’re going to be looking at performance, so set up timing to see how long things are taking. After each statement is run, the system will also provide a time output. Since this tutorial runs in a browser, there’s some variability in results.
\timing
There are two standard directions of tag queries:
- "What are the tags for this particular cat?"
- "What cats have this particular tag or set of tags?"
What Tags does this Cat have?
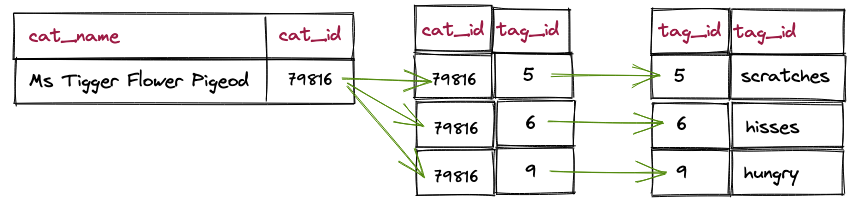
The query is simple, and the performance is very good (under 1 ms).
SELECT tag_name
FROM tags
JOIN cat_tags USING (tag_id)
WHERE cat_id = 444;
What cats have this tag?
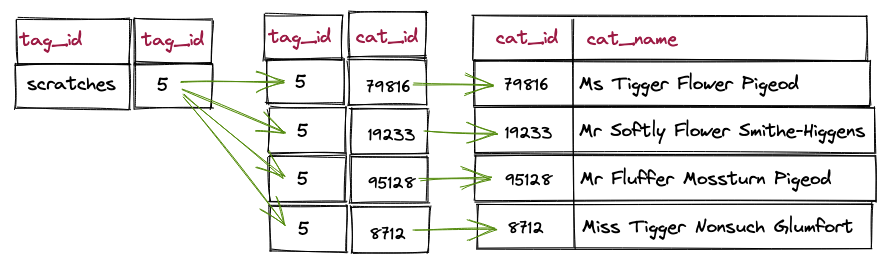
The query is still simple, and the performance is not unexpected for the number of records that meet the filter criterion.
SELECT Count(*)
FROM cats
JOIN cat_tags a ON (cats.cat_id = a.cat_id)
JOIN tags ta ON (a.tag_id = ta.tag_id)
WHERE ta.tag_name = 'brown';
What cats have these two tags?
This is where things start to come off the rails for the relational model, because finding just the records that have two particular tags involves quite complicated SQL.
SELECT Count(*)
FROM cats
JOIN cat_tags a ON (cats.cat_id = a.cat_id)
JOIN cat_tags b ON (a.cat_id = b.cat_id)
JOIN tags ta ON (a.tag_id = ta.tag_id)
JOIN tags tb ON (b.tag_id = tb.tag_id)
WHERE ta.tag_name = 'brown' AND tb.tag_name = 'aloof';
This query takes around >1s to find the cats that are both "brown" and "aloof".
What cats have these three tags?
Just so you can see the pattern, here's the three tag version.
SELECT Count(*)
FROM cats
JOIN cat_tags a ON (cats.cat_id = a.cat_id)
JOIN cat_tags b ON (a.cat_id = b.cat_id)
JOIN cat_tags c ON (b.cat_id = c.cat_id)
JOIN tags ta ON (a.tag_id = ta.tag_id)
JOIN tags tb ON (b.tag_id = tb.tag_id)
JOIN tags tc ON (c.tag_id = tc.tag_id)
WHERE ta.tag_name = 'brown'AND tb.tag_name = 'aloof'AND tc.tag_name = 'red';
At this point the decreasing number of records in the result set is balancing out the growing complexity of the multi-join and query time has only grown to >2s.
But imagine doing this for five, six or seven tags?
Tags in an integer array model
What if we changed our model, and instead of modeling the cat/tag relationship with a correlation table, we model it with an integer array?
CREATE TABLE cats_array (
cat_id serial primary key,
cat_name text not null,
cat_tags integer[]
);
Now our model has just two tables, cats_array and tags.
We can populate the array-based table from the relational data, so we can compare answers between models.
Data Generation SQL
INSERT INTO cats_array
SELECT cat_id, cat_name, array_agg(tag_id) AS cat_tags
FROM cats
JOIN cat_tags USING (cat_id)
GROUP BY cat_id, cat_name;
With this new data model, the size of the required tables has gone down, and we are using only 1040KB.
SELECT pg_size_pretty(
pg_total_relation_size('cats_array') +
pg_total_relation_size('tags'));
Once the data are loaded, we need the most important part -- an index on
the cat_tags integer array.
CREATE INDEX cats_array_x ON cats_array USING GIN (cat_tags);
This GIN index is a perfect fit for indexing collections (like our array) where there is a fixed and finite number of values in the collection (like our ten tags). While Postgres ships with an intarray extension, the core support for arrays and array indexes have caught up with and rendered much of the extension redundant.
As before, we will test common tag-based use cases.
What tags does this cat have?
The query is much less pretty! First we have to lookup the tag_id values
in cat_tags and use unnest() to expand them out into a relation. Then we're
ready to join that relation to the tags table to find the tag_name that
corresponds to the tag_id.
SELECT c.cat_id, c.cat_name, t.tag_name
FROM cats_array c
CROSS JOIN unnest(cat_tags) AS tag_id
JOIN tags t USING (tag_id)
WHERE cat_id = 779;
The query hits the cats primary key index and returns in the 100ms range.
Great performance!
What cats have these (three) tags?
This is the query that flummoxed our relational model. Let's jump straight to the hardest case and try to find all the cats that are "red", "brown" and "aloof".
WITH tags AS MATERIALIZED (
SELECT array_agg(tag_id) AS tag_ids
FROM tags
WHERE tag_name IN ('red', 'brown', 'aloof')
)
SELECT Count(*)
FROM cats_array
CROSS JOIN tags
WHERE cat_tags @> tags.tag_ids;
First we have to go into the tags table to make an array of tag_id entries
that correspond to our tags. Then we can use
the @> Postgres array operator to
test to see which cats have cat_tags arrays that contain the query array.
The query hits the GIN index on cat_tags and returns the count of cats in
around 200ms. About seven times faster than the same query on the relational
model!
Tags in a text array model
So if partially de-normalizing our data from a cats -> cat_tags -> tags model
to a cats -> tags model makes things faster... what if we went all the way to
the simplest model of all -- just cats?
CREATE TABLE cats_array_text (
cat_id serial primary key,
cat_name text not null,
cat_tags text[] not null
);
Again we can populate this new model directly from the relational model.
Data Generation SQL
INSERT INTO cats_array_text
SELECT cat_id, cat_name, array_agg(tag_name) AS cat_tags
FROM cats
JOIN cat_tags USING (cat_id)
JOIN tags USING (tag_id)
GROUP BY cat_id, cat_name;
The result is 1224KB, a little larger than the integer array version.
SELECT pg_size_pretty(
pg_total_relation_size('cats_array_text') +
pg_total_relation_size('tags'));
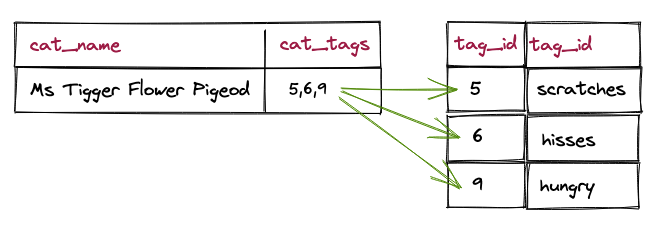
Now every cat has the tag names right in the record.
What tags does this cat have?
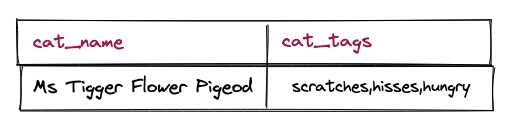
Since there's only one table with all the data, finding the tags of a cat is ridiculously easy.
SELECT cat_name, cat_tags
FROM cats_array_text
WHERE cat_id = 888;
What cats have these (three) tags?
Once again, in order to get good performance we need a GIN index on the array we will be searching.
CREATE INDEX cats_array_text_x ON cats_array_text USING GIN (cat_tags);
The query to find the cats are "red", "brown" and "aloof" is also wonderfully simple.
SELECT Count(*)
FROM cats_array_text
WHERE cat_tags @> ARRAY['red', 'brown', 'aloof'];
This query takes about the same amount of time as the integer array based query, 120ms.
Wins and losses
So are array-based models the final answer for tag-oriented query patterns in Postgres? On the plus side, the array models are:
- Faster to query;
- Smaller to store; and,
- Simpler to query!
Those are really compelling positives!
However, there are some caveats to keep in mind when working with these models:
- For the text-array model, there's no general place to lookup all tags. For a
list of all tags, you will have to scan the entire
catstable. - For the integer-array model, there's no way to create a simple constraint that
guarantees that integers used in the
cat_tagsinteger array exist in thetagstable. You can work around this with aTRIGGER, but it's not as clean as a relational foreign key constraint. - For both array models, the SQL can get a little crufty when the tags have to be un-nested to work with relational querying.